English
Privacy policy
Privacy Policy
Please carefully read the following policy (the "Privacy Policy") which describes how GenoScreen (the "Data Controller") collects, uses, discloses when necessary, stored, and otherwise manages personally identifiable information (the "Personal Data"). The Privacy Policy applies to all and any website, application, or service managed or offered by GenoScreen. GenoScreen may amend this policy from time to time by posting a revised version on its website, and without giving you notice, so please check this policy each time you visit the Site and note the date of publication. When you provide us with Personal Data in the manner described in Article 2 of this Privacy Policy, you allow us to collect it, store it and use it to: 1. Fulfill our contractual and business obligations towards you; 2. To administer, support, improve and develop our business in a legally justified manner (as explained in Article 3 of this Privacy Policy); All of this with being done with your consent, that you are allowed to withdraw anytime as described in Article 6 of this Privacy Policy. The EU General Data Protection Regulation (GDPR) replaces the Data Protection Directive 95/46/EC and was designed to harmonize data privacy laws across Europe, to protect and empower all EU citizens data privacy and to reshape the way organizations across the region approach data privacy. The key articles of the GDPR, as well as information on its business impact, can be found throughout this site: https://www.eugdpr.org/
Article 1 – Data Controller
The Data Controller of the Personal Data is GenoScreen, a French company with its headquarters situated at 1 Rue du Professeur Calmette – 59000 Lille, France, registered in the Company Register of Lille Métropole under the number 433996220.
Article 2 – Data collection
Subject to applicable laws, we collect information about you or any third party whose information you provide to us when you :
- Register to use our websites, applications or services (including free trials); this may include your name (or company name, if any), address, email address and phone number. We may also ask you to provide us with additional information about your business and your preferences;
- place an order using our websites, applications or services; this may include your name (or business name), your address and contact information (including phone number and email address) and payment;
- complete online forms (including reminder requests), take surveys, download information, such as white papers or other publications, or participate in any activity offered on the various interactive spaces present on our website or on our application or service;
- interact with us using social networks;
- provide your contact details when you register or access any website, application or service that we make available or when you update these details; and
- contact us offline, for instance by phone, fax, SMS, e-mail or mail.
We will also collect your Personal Data when you fill only a part of the fields to be filled on our site and / or on other online forms and / or when you abandon this entry along the way. We may use this data to contact you to remind you to enter missing information and / or for marketing purposes.
We may also collect information from your electronic devices (including mobile phones) and applications used by you or your users to access and use any of our websites, applications or services (for example, we may collect the identification number and the type of device used, geolocation and connection information, such as statistics on pages visited, traffic to and from sites, link referral URL, ad data, your IP address, your browsing history and your web log information). We will ask your authorization in advance before any step.
Subject to applicable law, we may supplement the Personal Data we collect with information obtained by third parties who are authorized to share it; for instance, information from credit institutions, providers seeking information or public sources (for instance, for the purpose of customer due diligence).
Third party data
If you provide us with Personal Data concerning a third party, it is your responsibility to ensure that you comply with applicable regulations regarding the protection of Personal Data and in particular with your obligations to obtain the prior consent of individuals. of which you provide us with the Personal Data. As such, in accordance with the applicable data protection regulations, you must have notified the data subject and obtained his express consent to provide us with his Personal Data and to have informed him of the manner in which we collect, use, disclose and retain Personal Data about him or invite him to read our Privacy Policy.
Article 3 – Use of Personal Data
Subject to applicable laws, we collect and process your Personal Data for the following purposes:
- provide any information and services you have requested and the applications or services you have ordered;
- compare the information for the purpose of checking its accuracy and corroborating it with third parties's data;
- provide, maintain, protect and enhance the applications, products, services and information you have requested from us;
- manage and administer the use you make of the applications, products and services you have asked us to provide;
- manage our business relationship (for instance, customer services and support activities);
- monitor, measure, enhance and protect our content, websites, applications and services and provide you with a personalized and quality user experience;
- perform, internally, controls on our websites, applications, systems and services to test and improve their security, provision and performance. Where applicable, we will use a pseudonymized form of any information used for such purposes, and we will ensure that such information is presented in a package and will not be binding on you or any other person involved;
- provide you with any information we are required to provide to you in order to comply with our regulatory or legal obligations;
- detect, investigate or prevent criminal, illegal or prohibited activities, or protect our rights (including liaising with law enforcement and regulatory agencies for these purposes);
- contact you to find out if you want to participate in our customer surveys (for example, feedback on your use of our applications, products and services);
- track and analyze statistics and comparisons jointly to avoid being identified or identifying any other individual;
- provide you with advertisements, marketing messages or targeted information that may be useful to you, based on your use of our applications and services;
- deliver content and services jointly with third parties with whom you have a separate relationship.
To the extent permitted by applicable law, we will retain your information after you cease to use GenoScreen websites, applications or services, for a limited time. This information will be kept and used for legal, regulatory, anti-fraud and legitimate conduct of our activities for the legally permitted duration.
Our websites, applications (including mobile apps) and services may contain technologies that allow us to:
- verify specific information from your device or systems necessary for your use of the websites, applications or services in relation to our records to ensure that the websites, applications or services are used in compliance with our Terms and Conditions and our agreements with the end user and for the purpose of solving potential problems;
- obtain information about technical errors or other issues related to our websites, applications and services;
- gather information about how you and the users make use of the features of our websites, applications and services.
You can manage your privacy settings in your browser or on our apps or services (if applicable).
GenoScreen works with MailChimp
GenoScreen sends all email communication through a viable third party named MailChimp. More than 12 million people and businesses around the world use MailChimp. MailChimp features and integrations allow their clients to send marketing emails, automated messages, and targeted campaigns.
Article 4 – Sharing of Personal Data
We may share your Personal Data with:
- our service providers and agents (including their subcontractors) or third parties who process information on our behalf (for instancee, Internet platform or service providers, payment processors and companies we use to help us send you communications) so that they can help us provide you with the applications, products, services and information you have been interested in receiving or which, from our point of view, would be likely to interest you;
- partners, including system builders, resellers, distributors, software vendors, and developers, who help us provide you with the applications, products, services, and information you've requested, or who our point of view, are likely to interest you;
- third parties that we have used for the execution of payment transactions, such as clearing companies, clearing systems, financial institutions and transaction beneficiaries;
- third parties, when you have a relationship with that third party and have consented to us transmitting information (for instance, social networking sites or other third-party application providers);
- third parties for marketing purposes (eg our partners and other third parties with whom we work and whose products, from our point of view, may be of interest to you in the conduct of your business activities. For instance: financial service providers (such as banks, insurers and financial service providers), payment solution providers, software providers and service providers that provide business solutions);
- credit reference and fraud prevention agencies;
- regulators, to meet the legal and regulatory obligations of GenoScreen;
- police authorities, so that they can detect or prevent crimes or prosecute offenders;
- any third party, in the context of existing or pending legal proceedings, provided we are legally entitled to do so (for example, in response to an order issued by a court);
- any third party to comply with our legal and regulatory obligations, including legal or regulatory reporting and the detection or prevention of unlawful acts;
- our own auditors and consultants, to fulfill our audit responsibilities;
- another company, if we sell or buy an activity or assets (or negotiate the sale or purchase of an activity or assets);
- any other company to which we may be liable to assign the contract which binds us; and
- public bodies that the law in force requires to inform.
We may share, publicly or with third parties, information about our websites, applications, products or services to the exclusion of any information that may identify you.
Article 5 – Marketing
We may occasionally use your information to contact you for the purpose of providing you with information about our applications, products and services that may be of interest to you. To this end, you can be contacted by phone, mail, SMS or e-mail. You have the right to ask us to stop contacting you for marketing purposes at any time. If you wish, you may exercise these rights by selecting your contact preferences where you provide us with your Personal Data on our websites, applications or services, using any preference center to which we give you access or in us, by sending an e-mail to This email address is being protected from spambots. You need JavaScript enabled to view it..
You can also unsubscribe from any marketing campaign and / or advertising by e-mail by clicking on the link included in the emails we send you.
Article 6 – Your rights
In some cases you will have the following rights (for more details, see https://www.eugdpr.org/):
- the right to know how we use your data and the right of access to your data;
- the right to request the modification or deletion of your data as well as to limit the processing of your data;
- the right to oppose the processing of your data, for example for marketing purposes or when their use is based on our legitimate interest;
- the right to receive data about you that you have provided automatically in a structured, commonly used and machine-readable format, or to have it sent directly to another company, if technically feasible ("Data Portability");
- when you have consented to the processing of your data, the right to withdraw your consent in accordance with legal or contractual limitations;
- the right to refuse any decision based on automated processing of your personal data, including profiling; and
- the right to lodge a complaint with the supervisory authority responsible for data protection issues (for instance, the National Commission for Data Protection and Freedoms).
If you request to receive a copy of your data, you may be required to pay certain fees.
If we have incorrect information about you or if your contact information has changed, we thank you for informing us so that we can correct and update our files.
If you withdraw your consent to the use of your personal data for the purposes set out in our Privacy Policy, we may, as a result, no longer be able to provide you with access to all or part of our websites, applications and services.
We will retain your personal data for the duration of our business relationship and after its expiration for the period of time necessary for the legitimate conduct of our business, or to comply with applicable laws and regulations. In cases where we no longer need your Personal Data, we will destroy it in a secure way (without sending you any other notice).
Article 7 - Storage and Processing
We ensure the security of your data by taking the necessary technical and structural measures to prevent their unlawful or unauthorized processing or accidental loss, destruction and / or damage. We strive to protect our Personal Data as much as we can. However, we can not guarantee the security of your data transmitted to our Internet sites, applications or services or to other Internet sites, applications and services via an Internet connection or any other connection. If we have given you (or if you have chosen) a password that allows you to access certain areas of our websites, applications or services, please keep it confidential; we will not share this password with anyone.
If you believe that your account has been hacked, please contact us at This email address is being protected from spambots. You need JavaScript enabled to view it.com.
Article 8 – Data transferred outside the EEA
In the European Union, Personal Data is protected by data protection laws. However, other countries do not necessarily protect your Personal Data in the same way.
Our websites, our services and applications may also be hosted in the United States or outside the EEA (which is made up of European Union as well as Norway, Iceland and Liechtenstein). This implies that we may transfer any data transmitted to us via the website, application or service outside the European Economic Area ("EEA") to the United States or Canada. other territories outside the EEA.
We may use the services of providers located outside the EEA to help us provide you with our websites, applications and services (such as, for example, platform or payment service providers who help us to deliver our services and applications or execute your payments). This means that we may transmit your data to service providers located outside the EEA for the purpose of providing you with our applications and services.
We make sure that when your service providers and hosting providers transfer your data outside the EEA, appropriate measures and controls are implemented in compliance with applicable laws and regulations to protect them. In each case, these transfers are made in accordance with the requirements of Regulation (EU) 2016/679 (General Data Protection Regulation) and may be based on the use of standard clauses on transfers of personal data outside the EEA of the European Commission.
By using our websites, products and services or interacting with us under the conditions described in this Privacy Policy, you consent to the transfer of your data outside the EEA in the cases provided for in this Privacy Policy. If you refuse to transfer your data outside the EEA, you will not be able to use our websites, applications or services.
Google Analytics uses "cookies" to enable the websites to analyze how users use the websites, applications and services. Information about your use of the Internet sites, applications or services (including your IP address) generated by the cookie are transmitted to Google for storage on servers located in the United States. Google will use this information to evaluate your use of websites, applications and services, to compile reports on website activity for site operators and to provide other services related to the activity of the website and the use of the Internet. Google may also disclose this information to third parties when required by law or when such third parties process the information on Google's behalf. Google will not associate your IP address with other data held by Google. You can refuse the use of cookies by selecting the appropriate settings in your browser or application. Note however that your refusal may result in partial unavailability of the website. For more information, refer to "How Google Uses Certain Data Collected When You Use Partner Sites or Applications" (available at www.google.com/policies/privacy/partners/ or any other link that Google may provide to you).
If you want Google Analytics to stop following you on all websites, visit http://tools.google.com/dlpage/gaoptout.
Article 10 – Questions or comments
If you have questions or comments about this privacy policy, please email us at This email address is being protected from spambots. You need JavaScript enabled to view it.com or write to GenoScreen, 1 Rue du Professeur Calmette, 59000 Lille - France.
console.log('hello');
Publications
Publications with GenoScreen researchers in the authors
(79 publications from 2008 to 2023)
-
Yenew B., Ghodousi A., …, Gaudin C., …, Supply P. et al., A smooth tubercle bacillus from Ethiopia phylogenetically close to the Mycobacterium tuberculosis complex, Nat. Commun., Nov. 2023, 14(1):7519. doi: 10.1038/s41467-023-42755-9.
- Jouet A, Braet SM, Gaudin C, Bisch G, Vasconcellos S, Epaminondas Nicacio de Oliveira do Livramento RE, Prado Palacios YY, Fontes AB, Lucena N, Rosa P, Moraes M, La K, Badalato N, Lenoir E, Ferré A, Clément M, Hasker E, Grillone SH, Abdou W, Said A, Assoumani Y, Attoumani N, Laurent Y, Cambau E, de Jong BC, Suffys PN, Supply P. Hi-plex deep amplicon sequencing for identification, high-resolution genotyping and multidrug resistance prediction of Mycobacterium leprae directly from patient biopsies by using Deeplex Myc-Lep. EBioMedicine. 2023 Jul;93:104649. doi: 10.1016/j.ebiom.2023.104649. Epub 2023 Jun 14. PMID: 37327675; PMCID: PMC10279541.
- Vallier M., Segurens B., Larsonneur E., Meyer V., Ferreira S., Christophe Caloustian C., Jean-François Deleuze JF., Dougados M., Chamaillard M., Characterisation of gut microbiota composition in patients with axial spondyloarthritis and its modulation by TNF inhibitor treatment. RMD Open, March 2023, 9:e002794. doi: 10.1136/rmdopen-2022-002794.
- Merker M, Rasigade JP, Barbier M, Cox H, Feuerriegel S, A. Kohl T, Shitikov E, Klaos K, Gaudin C, Antoine R, Diel R, Borrell S, Gagneux S, Nikolayevskyy V, Andres S, Crudu V, Supply P, Niemann S, Wirth T. Transcontinental spread and evolution of Mycobacterium tuberculosis W148 European/Russian clade toward extensively drug resistant tuberculosis. Nature Communications. 2021-2022 Oct-Aug
- Marijke Braet S, Jouet A, Aubry A, Van Dyck-Lippens M, Lenoir E, Assoumani Y, Baco A, Mzembaba A, Cambau E, Ezidio Gonçalves Vasconcellos S, Rigouts L, Suffys PN, Hasker E, Supply P, de Jong BC. Investigating drug resistance of Mycobacterium leprae in the Comoros: an observational deep-sequencing study. Lancet. 2022 July.
- Camille d'Humières C , Nadia Gaïa N., …. Stéphanie Ferreira et al, Contribution of Clinical Metagenomics to the Diagnosis of Bone and Joint Infections, Front. Microbiol. 2022 Apr 21; 13:863777.
- Sokol H, Contreras V, Maisonnasse P, Desmons A, Delache B, Sencio V, Machelart A, Brisebarre A, Humbert L, Deryuter L, Gauliard E, Heumel S, Rainteau D, Dereuddre-Bosquet N, Menu E, Ho Tsong Fang R, Lamaziere A, Brot L, Wahl C, Oeuvray C, Rolhion N, Van Der Werf S, Ferreira S, Le Grand R, Trottein F. SARS-CoV-2 infection in nonhuman primates alters the composition and functional activity of the gut microbiota. Gut Microbes. 2021 Jan-Dec, 13(1):1-19
- Hilaire P, Landemaine L, Contreras S, Blanquart-Goudezeune H, Siguier P, Cornet F, Chiapello H, Loux V, Clavaud C, Morand SC. Complete Genome Sequence of Staphylococcus epidermidis PH1-28, Isolated from the Forehead of a Hyperseborrheic Donor. Microbiol Resour Announc. 2021 March, 10(9):e00165-21
- Allen AC, Malaga W, Gaudin C, Volle A, Moreau F, Hassan A, Astarie-Dequeker C, Peixoto A, Antoine R, Pawlik A, Frigui W, Berrone C, Brosch R, Supply P, Guilhot C. Parallel in vivo experimental evolution reveals that increased stress resistance was key for the emergence of persistent tuberculosis bacilli. Nat Microbiol. 2021 Aug, 6(8):1082-1093
- Hilaire P, Contreras S, Blanquart-Goudezeune H, Verbeke J, Delépine B, Marmiesse L, Peyraud R, Morand SC. Complete Genome Sequence of Sphingobium xenophagum PH3-15, Isolated from La Roche-Posay Thermal Water Sources. Microbiol Resour Announc. 2021 Aug, 10(33):e0070021
- Hellal J., Joulian C., Urien C., Ferreira S., Denonfoux J., Hermon L., Vuilleumier S., Imfeld G., Chlorinated ethene biodegradation and associated bacterial taxa in multi-polluted groundwater: Insights from biomolecular markers and stable isotope analysis, Science of The Total Environment, 2021 apr; 763:142950
- Jouet A, Gaudin C, Badalato N, Allix-Béguec C, Duthoy S, Ferré A, Diels M, Laurent Y, Contreras S, Feuerriegel S, Niemann S, André E, Kaswa MK, Tagliani E, Cabibbe A, Mathys V, Cirillo D, de Jong BC, Rigouts L, Supply P. Deep amplicon sequencing for culture-free prediction of susceptibility or resistance to 13 anti-tuberculous drugs. Eur Respir J. 2020 Sep 17:2002338.
- Vandenborght LE, Enaud R, Urien C, Coron N, Girodet PO, Ferreira S, Berger P, Delhaes L. Type 2-high asthma is associated with a specific indoor mycobiome and microbiome. J Allergy Clin Immunol. 2020 Sep 11:S0091-6749(20)31248-3
- Feuerriegel S, Kohl TA, Utpatel C, Andres S, Maurer FP, Heyckendorf J, Jouet A, Badalato N, Foray L, Kamara RF, Conteh OS, Supply P, Niemann S. Rapid genomic first- and second-line drug resistance prediction from clinical Mycobacterium tuberculosis specimens using Deeplex®-MycTB. Eur Respir J. 2020 Aug 6:2001796.
- Foligné B, George F, Standaert A, Garat A, Poiret S, Peucelle V, Ferreira S, Sobry H, Muharram G, Lucau-Danila A, Daniel C. High-dose dietary supplementation with zinc prevents gut inflammation: Investigation of the role of metallothioneins and beyond by transcriptomic and metagenomic studies. FASEB J. 2020 Jul 30.
- Ngabonziza JCS, Loiseau C, Marceau M, Jouet A, Menardo F, Tzfadia O, Antoine R, Niyigena EB, Mulders W, Fissette K, Diels M, Gaudin C, Duthoy S, Ssengooba W, André E, Kaswa MK, Habimana YM, Brites D, Affolabi D, Mazarati JB, de Jong BC, Rigouts L, Gagneux S, Meehan CJ, Supply P. A sister lineage of the Mycobacterium tuberculosis complex discovered in the African Great Lakes region. Nat Commun. 2020 Jun 9;11(1):2917.
- Sencio V, Barthelemy A, Tavares LP, Machado MG, Soulard D, Cuinat C, Queiroz-Junior CM, Noordine ML, Salomé-Desnoulez S, Deryuter L, Foligné B, Wahl C, Frisch B, Vieira AT, Paget C, Milligan G, Ulven T, Wolowczuk I, Faveeuw C, Le Goffic R, Thomas M, Ferreira S, Teixeira MM, Trottein F. Gut Dysbiosis during Influenza Contributes to Pulmonary Pneumococcal Superinfection through Altered Short-Chain Fatty Acid Production. Cell Rep. 2020 Mar 3;30(9):2934-2947.
- Soret P, Vandenborght LE, Francis F, Coron N, Enaud R, Avalos M, Schaeverbeke T, Berger P, Fayon M, Thiebaut R, Delhaes L; Mucofong Investigation Group. Respiratory mycobiome and suggestion of inter-kingdom network during acute pulmonary exacerbation in cystic fibrosis. Sci Rep. 2020 Feb 27;10(1):3589.
- El Achkar S, Demanche C, Osman M, Rafei R, Ismail MB, Gaudin C, Duthoy S, De Matos F, Yaacoub H, Pinçon C, Hamze M, Supply P. Zoonotic tuberculosis in humans assessed by next-generation sequencing: an 18-month nationwide study in Lebanon. Eur Respir J. 2020 Jan 2;55(1).
- Saint-Lu N, Burdet C, Sablier-Gallis F, Corbel T, Nevière A, Sayah-Jeanne S, Pulse M, Weiss W, Ferreira S, Andremont A, Mentré F, de Gunzburg J. DAV131A Protects Hamsters from Lethal Clostridioides difficile Infection Induced by Fluoroquinolones. Antimicrob Agents Chemother. 2019 Dec 20;64(1)
- Ng KCS, Supply P, Cobelens FGJ, Gaudin C, Gonzalez-Martin J, de Jong BC, Rigouts L. How well do routine molecular diagnostics detect rifampicin heteroresistance in Mycobacterium tuberculosis? J Clin Microbiol. 2019 Aug 14.
- Burdet C, Nguyen TT, Duval X, Ferreira S, Andremont A, Guedj J, Mentré F; DAV132-CL-1002 study group. Impact of antibiotic gut exposure on the temporal changes in microbiome diversity. Antimicrob Agents Chemother. 2019 Jul 15.
- Hermon L, Hellal J, Denonfoux J, Vuilleumier S, Imfeld G, Urien C, Ferreira S, Joulian C. Functional Genes and Bacterial Communities During Organohalide Respiration of Chloroethenes in Microcosms of Multi-Contaminated Groundwater. Front Microbiol. 2019 Feb 12;10:89.
- Pedersen MK, Andersen AB, Folkvardsen DB, Rasmussen EM, Svensson E, Lillebaek T, Supply P. Set-up and validation of mycobacterial interspersed repetitive unit-variable number of tandem repeat (MIRU-VNTR) analysis of Mycobacterium tuberculosis using BioNumerics software. PLoS. 2018 Oct 31;13(10).
- Makhado NA, Matabane E, Faccin M, Pinçon C, Jouet A, Boutachkourt F, Goeminne L, Gaudin C, Maphalala G, Beckert P, Niemann S, Delvenne JC, Delmée M, Razwiedani L, Nchabeleng M, Supply P, de Jong BC, André E.Outbreak of multidrug-resistant tuberculosis in South Africa undetected by WHO-endorsed commercial tests: an observational study.Lancet Infect Dis. 2018 Dec;18(12):1350-1359.
- Burdet C, Sayah-Jeanne S, Nguyen TT, Hugon P, Sablier-Gallis F, Saint-Lu N, Corbel T, Ferreira S, Pulse M, Weiss W, Andremont A, Mentré F, de Gunzburg J. Antibiotic-induced dysbiosis predicts mortality in an animal model of Clostridium difficile infection. Antimicrob Agents Chemother. 2018 Jul 30.
- Hermon L, Denonfoux J, Hellal J, Joulian C, Ferreira S, Vuilleumier S, Imfeld G. Dichloromethane biodegradation in multi-contaminated groundwater: Insights from biomolecular and compound-specific isotope analyses.Water Res. 2018 Oct 1;142:217-226.
- Contreras S, Sagory-Zalkind P, Blanquart H, Iltis A, Morand S. Complete Genome Sequence of Vitreoscilla filiformis (ATCC 15551), Used as a Cosmetic Ingredient. Genome Announc. 2017 Aug 24;5(34)
- Clément Y, Sarah G, Holtz Y, Homa F, Pointet S, Contreras S, Nabholz B, Sabot F, Sauné L, Ardisson M, Bacilieri R, Besnard G, Berger A, Cardi C, De Bellis F, Fouet O, Jourda C, Khadari B, Lanaud C, Leroy T, Pot D, Sauvage C, Scarcelli N, Tregear J, Vigouroux Y, Yahiaoui N, Ruiz M, Santoni S, Labouisse JP, Pham JL, David J, Glémin S. Evolutionary forces affecting synonymous variations in plant genomes. PLoS , 2017; May.22, volume 13
- Bruffaerts N, Vluggen C, Roupie V, Duytschaever L, Van den Poel C, Denoël J, Wattiez R, Letesson JJ, Fretin D, Rigouts L, Chapeira O, Mathys V, Saegerman C, Huygen K. Virulence and immunogenicity of genetically defined human and porcine isolates of M. avium subsp. hominissuis in an experimental mouse infection. PLoS , 2017; Feb.9, volume 12
- Dugat-Bony E, Garnier L, Denonfoux J, Ferreira S, Sarthou AS, Bonnarme P, Irlinger F4. Highlighting the microbial diversity of 12 French cheese varieties. International Journal of Food Microbiology, Volume 238, 5 December 2016, Pages 265-273.
- Bruffaerts N, Vluggen C, Duytschaever L, Mathys V, Saegerman C, Chapeira O, Huygen K. Genome Sequences of Four Strains of Mycobacterium avium subsp. hominissuis, Isolated from Swine and Humans, Differing in Virulence in a Murine Intranasal Infection Model. Genome Announc. 2016; June 16; vol 4, issue 3.
- Ribière C, Beugnot R, Parisot N, Gasc C, Defois C, Denonfoux J, Boucher D, Peyretaillade E, Peyret P. Targeted Gene Capture by Hybridization to Illuminate Ecosystem Functioning. Methods Mol Biol. 2016; 1399:167-82
- Parisot N, Peyretaillade E, Dugat-Bony E, Denonfoux J, Mahul A, Peyret P. Probe Design Strategies for Oligonucleotide Microarrays. Methods Mol Biol. 2016; 1368:67-82
- T. Bouchez, A. L. Blieux, S. Dequiedt, I. Domaizon, A. Dufresne, S. Ferreira, J. J. Godon, J. Hellal, C. Joulian, A. Quaiser5 • F. Martin-Laurent, A. Mauffret, J. M. Monier, P. Peyret, P. Schmitt-Koplin, O. Sibourg, E. D’oiron, A. Bispo, I. Deportes, C. Grand, P. Cuny, P. A. Maron, L. Ranjard. Molecular microbiology methods for environmental diagnosis. Environ Chem Lett (2016), vol 14, issu 4, pages 423-441.
- Plé C, Vandenborght LE, Adele-Dit-Renseville N, Breton J, Daniel C, Ferreira S, Benoît F. Snapshot on a Pilot Metagenomic Study for the Appraisal of Gut Microbial Diversity in Mice, Cat, and Man. Gastroenterol Res Pract. 2016; 2016:6587825. Epub 2016 Apr 24.
- S. Roisin, C. Gaudin, R. De Mendonça, J. Bellon, K. Van Vaerenbergh, K. De Bruyne, B. Byl, H. Pouseele, O. Denis, P. Supply. Pan-genome multilocus sequence typing and outbreak-specific reference-based single nucleotide polymorphism analysis to resolve two concurrent Staphylococcus aureus outbreaks in neonatal services Clinical Microbiology and Infection, Volume 22, Issue 6, June 2016, Pages 520-526.
- Pankhurst LJ, Del Ojo Elias C, Votintseva AA, Walker TM, Cole K, Davies J, Fermont JM, Gascoyne-Binzi DM, Kohl TA, Kong C, Lemaitre N, Niemann S, Paul J, Rogers TR, Roycroft E, Smith EG, Supply P, Tang P, Wilcox MH, Wordsworth S, Wyllie D, Xu L, Crook DW. Rapid, comprehensive, and affordable mycobacterial diagnosis with whole-genome sequencing: a prospective study. The Lancet Respiratory Medicine, Volume 4, Issue 1, January 2016, Pages 49-58
- Annales de Dermatologie et de Vénéréologie, Volume 142, Issue 12, Supplement, December 2015, Pages S617
- PLoS . 2015 Nov 6;10(11)
- Journal of Functional Foods, Volume 18, Part A, October 2015, Pages 575-585
- The Lancet Infectious Diseases, Volume 15, Issue 10, October 2015, Pages 1193-1202
- Lancet Respir Med.- 2015 Dec 3
- FEMS Microbiol Ecol. 2015 May;91(5)
- Genomics Data, Volume 4, June 2015, Pages 22-23
- Nat Genet. 2015 Mar;47(3):242-9
- Microb Biotechnol - 2015 Jan;8(1):131-42
- Genom Data. 2015 Feb 2;4:22-3
- PLoS . 2015 Jan 13;10
- PLoS . 2014 Oct 14;9
- J Clin Microbiol. 2014 Sep;52(9)
- Methods in Microbiology, Volume 42, 2015, Pages 359-394
- Science of the Total Environment, Volumes 485–486, 1 July 2014, Pages 508-517
- Neurology. 2014 Mar 25;82(12)
- Indoor Air. 2014 Feb;24(1):29-40
- BMC Genomics. 2014 Apr 8;15(1):272
- Int J Tuberc Lung Dis 18:594-600
- Clinical Microbiology and Infection, Volume 20, Issue 1, January 2014, Pages 70-78
- J Clin Microbiol. 2014 Jan;52(1):164-72
- Current Opinion in Biotechnology, Volume 24, Supplement 1, July 2013, Page S37
- J Gen Virol. 2013 December.
- PLoS . 2013 Oct 17; 8(10): e77635.
- Sex Trans Infect: Volume 40, Number 8, August 2013.
- J Clin Microbiol. 2013 Jul; 51(7):2427-3
- Indoor Air. 2013 May 27.
- PLoS . 2013 May 9; 8(5):e63128.
- Clin Microbiol Infect. 2014 Jan;20(1):70-8.
- PLoS . 2013 Feb 1; 8(2): e55598.
- J Bacteriol. 2012 Apr;194(7):1838-9.
- PLoS . 2012;7(4):e36313. Epub 2012 Apr 27.
- J Clin Microbiol. 2012 May;50(5):1830-1.
- Mol Ecol Resour. 2011 Jul; 11(4):638-44.
- BMC Genomics. 2011 Apr 27;12:207.
- PLoS . 2011 Mar 25;6(3):e18256.
- BMC Genomics, 12:245, 2011 May 19
- JCM 2011,10.1128, December 2011
- Respiratory Medicine, Volume 105, Supplement 1, October 2011, Pages S67-S73
- Rev Port Pneumol. 2010 Jan;16SA:S89-93.
- Revista Portuguesa de Pneumologia, Volume 16, Supplement A, January 2010, Pages S89-S93
- Molecular Ecology Resources, 2010 Dec
- BMC Genomics. 2010 Oct 12; 11:560.
- J Clin Microbiol. 2008 Aug;46(8):2692-9.
Publications with GenoScreen quoted in the text (1635 publications from 2008 to 2024)
GenoScreen is quoted in the text or in acknowledgements we find: "GenoScreen"
-
Schwarz J.M. et al., Diverse pollen nutrition can improve the development of solitary bees but does not mitigate negative pesticide impacts, Science of The Total Environment, Volume 912, 20 February 2024, 169494.
-
Vautrin et al., The short-term response of soil microbial communities to digestate application depends on the characteristics of the digestate and soil type, Applied Soil Ecology, Volume 193, January 2024, 105105.
-
Priyanka Guha, Siddhartha Dutta, Krishna Murti, Jay Karan Charan, Krishna Pandey, V. Ravichandiran, Sameer Dhingra. The Integration of Omics: A Promising Approach to Personalized Tuberculosis Treatment. Medicine in Omics. 2024, 100033, https://doi.org/10.1016/j.meomic.2024.100033
-
Laura Niebling, Ramona Nitzsche, Thorben Sieksmeyer, Vera Haskamp, Jonas Kissenkötter, Ahmed Abd El Wahed, Thomas Teufel, Herbert Hermann, Nils Paust and Ana R. Homann. Fast and on-site animal species identification in processed meat via centrifugal microfluidics and isothermal amplification. Lab Chip. 2024. DOI: 10.1039/D3LC01103.
-
Rodrigues, C., Singhal, T. What is New in the Diagnosis of Childhood Tuberculosis?. Indian J Pediatr (2024). https://doi.org/10.1007/s12098-023-04992-0
-
Gupta, R.K., Anthwal, D., Bhalla, M. et al. Direct Detection of Fluoroquinolone Resistance in Sputum Samples from Tuberculosis Patients by High Resolution Melt Curve Analysis. Curr Microbiol. 81, 27 (2024). https://doi.org/10.1007/s00284-023-03519-2
-
Vallier M, Segurens B, Larsonneur E, Meyer V, Ferreira S, Caloustian C, Deleuze JF, Dougados M, Chamaillard M, Miceli-Richard C. Characterisation of gut microbiota composition in patients with axial spondyloarthritis and its modulation by TNF inhibitor treatment. RMD Open. 2023 Mar;9(1):e002794. doi: 10.1136/rmdopen-2022-002794.
- Seanne P. Buckwalter, Sara L. Olson, Madiha Fida, L. Elaine Epperson, Nabeeh A. Hasan, Reeti Khare, Michael Strong, Nancy L. Wengenack. Mycobacterium abscessus subspecies identification using the Deeplex Myc-TB targeted NGS assay. Journal of Clinical Microbiology. 2023 Sept. doi: https://doi.org/10.1128/jcm.00489-23.
- Mansoor H, Hirani N, Chavan V, Das M, Sharma J, Bharati M, Oswal V, Iyer A, Morales M, Joshi A, Ferlazzo G, Isaakidis P, Ndlovu Z, England K. Clinical utility of target-based next-generation sequencing for drug-resistant TB. Int J Tuberc Lung Dis. 2023 Jan 1;27(1):41-48. doi: 10.5588/ijtld.22.0138.
- Murphy SG, Smith C, Lapierre P, Shea J, Patel K, Halse TA, Dickinson M, Escuyer V, Rowlinson MC, Musser KA. Direct detection of drug-resistant Mycobacterium tuberculosis using targeted next generation sequencing. Front Public Health. 2023 Jun 29;11:1206056. doi: 10.3389/fpubh.2023.1206056.
- Iyer A, Ndlovu Z, Sharma J, Mansoor H, Bharati M, Kolan S, Morales M, Das M, Issakidis P, Ferlazzo G, Hirani N, Joshi A, Tipre P, Sutar N, England K. Operationalising targeted next-generation sequencing for routine diagnosis of drug-resistant TB. Public Health Action. 2023 Jun 21;13(2):43-49. doi: 10.5588/pha.22.0041.
- Kuroiwa, K., Danilo, B., Perrot, L., Thenault, C., Veillet, F., Delacote, F., Duchateau, P., Nogué, F., Mazier, M. and Gallois, J.-L. (2023), An iterative gene-editing strategy broadens eIF4E1 genetic diversity in Solanum lycopersicum and generates resistance to multiple potyvirus isolates. Plant Biotechnol. J, 21: 918-930. doi: https://doi.org/10.1111/pbi.14003.
- Kiyoharu Fukushima, Yuki Matsumoto, Takanori Matsuki, Haruko Saito, Daisuke Motooka, Sho Komukai, Eriko Fukui. MGIT-seq for the Identification of Nontuberculous Mycobacteria and Drug Resistance: a Prospective Study. Journal of Clinical Microbiology. 2023 Mar;61:4. https://doi.org/10.1128/jcm.01626-22.
- Murphy Shannon G., Smith Carol, Lapierre Pascal, Shea Joseph, Patel Kruthikaben, Halse Tanya A., Dickinson Michelle, Escuyer Vincent, Rowlinson Marie Claire, Musser Kimberlee A. Direct detection of drug-resistant Mycobacterium tuberculosis using targeted next generation sequencing. Frontiers in Public Health. 2023 Sept. 11. doi: 10.3389/fpubh.2023.1206056.
- Charles C, Conde C, Vorimore F, Cochard T, Michelet L, Boschiroli ML, Biet F. Features of Mycobacterium bovis Complete Genomes Belonging to 5 Different Lineages. Microorganisms. 2023; 11(1):177. doi: https://doi.org/10.3390/microorganisms11010177.
- Callejon S, Giraud F, Larue F, Buisson A, Mateos L, Grare L, Guyoux A, Perrier E, Ardiet N, Trompezinski S. Impact of Leave-on Skin Care Products on the Preservation of Skin Microbiome: An Exploration of Ecobiological Approach. Clin Cosmet Investig Dermatol. 2023 Sep 29;16:2727-2735. doi: 10.2147/CCID.S409583.
- Landemaine Leslie, Da Costa Gregory, Fissier Elsa, Francis Carine, Morand Stanislas, Verbeke Jonathan, Michel Marie-Laure, Briandet Romain, Sokol Harry, Gueniche Audrey, Bernard Dominique, Chatel Jean-Marc, Aguilar Luc, Langella Philippe, Clavaud Cecile, Richard Mathias L. Staphylococcus epidermidis isolates from atopic or healthy skin have opposite effect on skin cells: potential implication of the AHR pathway modulation. Frontiers in Immunology. 2023 May;14. doi: 10.3389/fimmu.2023.1098160.
- Le Loarer A, Marcellin-Gros R, Dufossé L, Bignon J, Frédérich M, Ledoux A, Queiroz EF, Wolfender J-L, Gauvin-Bialecki A, Fouillaud M. Prioritization of Microorganisms Isolated from the Indian Ocean Sponge Scopalina hapalia Based on Metabolomic Diversity and Biological Activity for the Discovery of Natural Products. Microorganisms. 2023; 11(3):697. https://doi.org/10.3390/microorganisms11030697.
- European Alzheimer’s & Dementia Biobank Mendelian Randomization (EADB-MR) Collaboration. Genetic Associations Between Modifiable Risk Factors and Alzheimer Disease. JAMA Netw Open. 2023;6(5):e2313734. doi:10.1001/jamanetworkopen.2023.13734
- Verneau O, Melliti S, Kimdil L, El Mouden EH, Achouri MS, Rouag R. Molecular Phylogenies of Leeches and Haemoparasites Infecting Freshwater Turtles in Aquatic Ecosystems of Northern Africa Suggest Phylogenetic Congruence between Placobdella costata Sensu Lato and Haemogregarina stepanowi Sensu Lato. Microorganisms. 2023; 11(6):1584. https://doi.org/10.3390/microorganisms11061584
- Callejon S, Giraud F, Larue F, Buisson A, Mateos L, Grare L, Guyoux A, Perrier E, Ardiet N, Trompezinski S. Impact of Leave-on Skin Care Products on the Preservation of Skin Microbiome: An Exploration of Ecobiological Approach. Clin Cosmet Investig Dermatol. 2023;16:2727-2735. https://doi.org/10.2147/CCID.S409583
- C. De Moya‐Ruiz, M. Juárez, P. Gómez, First report of Pepo aphid‐borne yellows virus on watermelon plants in Spain. New Disease Reports. 10.1002/ndr2.12215, 48, 1, (2023). doi: https://doi.org/10.1002/ndr2.124
- Camille Lebarbenchon, Steven M Goodman, Axel O G Hoarau, Gildas Le Minter, Andréa dos Santos, et al.. Bombali Ebolavirus in Mops condylurus Bats (Molossidae), Mozambique. Emerging Infectious Diseases. 2022, 28 (12), pp.2583 - 2585.
- Etc.
-
Amira C. et al, Revision of the systematics of the Polystomoidinae (Platyhelminthes, Monogenea, Polystomatidae) with redefinition of Polystomoides Ward, 1917 and Uteropolystomoides Tinsley, 2017. Parasite. 2022 December 16(29)56 :19.
-
Domitille J. et al, High-quality genome of the basidiomycete yeast Dioszegia hungarica PDD-24b-2 isolated from cloud water. G3 Genes|Genomes|Genetics. 2022 December (12)12.
-
Tara E. Ness et al, Optimizing DNA Extraction from Pediatric Stool for Diagnosis of Tuberculosis and Use in Next-Generation Sequencing Applications. ASM Journal. 2022 December 8(11) ;1.
-
Post R.J. et al, Identification of the onchocerciasis vector in the Kakoi-Koda focus of the Democratic Republic of Congo. PLOS Neglected Tropical Diseases. 2022 November 4(11);e0010684.
-
Antsa R. et al, MALDI-TOF MS: An effective tool for a global surveillance of dengue vector species. PLoS ONE. 2022 October 20(10)e0276488.
-
Philip W. Fowler et al, Epidemiological cut-off values for a 96-well broth microdilution plate for high-throughput research antibiotic susceptibility testing of M. tuberculosis. European Respiratory Journal. 2022 October(4) ;60 : 2200239.
-
Mishra et al, Stroke genetics informs drug discovery and risk prediction across ancestries. Nature. 2022 September 30(611) ;115-123.
-
Annelies Van Rie et al, Balancing access to BPaLM regimens and risk of resistance. The Lancet Infectious Diseases. 2022 August 22(22) ;10 :1411-1412.
-
Alffenaar JWC et al, Clinical standards for the dosing and management of TB drugs. The international journal of tuberculosis and lung disease : the official journal of the International Union against Tuberculosis and Lung Disease. 2022 June 1(6) ;26:483-499.
-
Doctor B. Sibandze et al, Rapid molecular diagnostics of tuberculosis resistance by targeted stool sequencing. Genome Medicine. 2022 May 19(14) ;52 :3392.
-
Edoardo M. et al, Ectomycorrhizal fungal communities of Swiss stone pine (Pinus cembra) depend on climate and tree age in natural forests of the Alps. Plant and Soil. 2022 May 26 ;734.
-
Yann L.G. et al, Association of Rare APOE Missense Variants V236E and R251G With Risk of Alzheimer Disease. Jama Neurol. 2022 May 31(79) ;7:652-663.
-
Günther G. et al, Drug-resistant tuberculosis: advances in diagnosis and management. Current Opinion in Pulmonary Medicine. 2022 May(3) ;28 :211-217.
-
Sophia B. Georghiou et al, Updating the approaches to define susceptibility and resistance to anti-tuberculosis agents: implications for diagnosis and treatment. Eur RespirJ. 2022 March. 59: 2200166.
- Radha Gopalaswamy et al, An odyssey from laboratory to field ? – Portable tNGS system for TB diagnosis in programmatic setting, Indian Journal of Tuberculosis, 2022 april, In Press
- Bourahima Kone et al, Molecular epidemiology and genetic diversity of Mycobacterium tuberculosis complex in referral health centers of Bamako, Mali: What is new?, International Journal of Infectious Diseases, 2022 april, 117:204-211
- Emmanuelle Boix et al, Synergistic interaction between pH and NaCl in the limits of germination and outgrowth of Clostridium sporogenes and Group I Clostridium botulinum vegetative cells and spores after heat treatment, Food Microbiology, 2022 april, In Press 104055
- Alain Din Dipita et al, DNA-typing improves illegal wildlife trade surveys: Tracing the Cameroonian bushmeat trade, Biological Conservation, 2022 may, 269:109552
- Michelle Li Wei Kam et al, Country-wide genotyping of Mycobacterium tuberculosis complex in Singapore, 2011–2017, Tuberculosis, 2022 may, 134:102204
- Kyoka Kuroiwa et al, CRISPR-based knock-out of eIF4E2 in a cherry tomato background successfully recapitulates resistance to pepper veinal mottle virus, Plant Science, 2022 march, 316:111160
- Anna S Dean et al, 25 years of surveillance of drug-resistant tuberculosis: achievements, challenges, and way forward, The Lancet Infectious Diseases, 2022 march, In Press
- Charlotte Amy et al, Are native phosphate solubilizing bacteria a relevant alternative to mineral fertilizations for crops? Part I. when rhizobacteria meet plant P requirements, Rhizosphere, 2022 march, 21:100476
- Pierre-François Perroud et al, Prime Editing in the model plant Physcomitrium patens and its potential in the tetraploid potato, Plant Science, 2022 march, 316:111162
- Simon Tiberi et al, Drug resistant TB – latest developments in epidemiology, diagnostics and management, International Journal of Infectious Diseases, 2022 march, In Press
- Elisabeth Hodille et al, Molecular tests for tuberculosis diagnosis : a major step forward to tackle the pandemics, Médecine et Maladies Infectieuses Formation, 2022 jan., In Press
- Stéphane Hourdez et al, Investigation of Capitella spp. symbionts in the context of varying anthropic pressures: First occurrence of a transient advantageous epibiosis with the giant bacteria Thiomargarita sp. to survive seasonal increases of sulfides in sediments, Science of The Total Environment, 2021 dec., 798:149149
- Malika Oubohssaine et al, The rhizosphere of Sulla spinosissima growing in abandoned mining soils is a reservoir of heavy metals tolerant plant growth-promoting rhizobacteria, Biocatalysis and Agricultural Biotechnology, 2021 dec., 102236Jun Niimi et al, Aroma and bacterial communities dramatically change with storage of fresh white truffle Tuber magnatum, LWT, 2021 nov., 151:112125
- Stéphanie Eyssautier-Chuine et al, Comparison of biofilm development on three building and restoration stones used in French monuments, International Biodeterioration & Biodegradation, 2021 nov., 165:105322
- Cyrine REZGUI et al, Linking changes in the soil microbial community to C and N dynamics during crop residue decomposition, Journal of Integrative Agriculture, 2021 nov., 20(11):3039-3059
- Christian M.J. Delannoy et al, Streptococcus agalactiae serotype IV in farmed tilapia, Aquaculture, 2021 nov., 544:737033
- Isabelle Bonnet et al, A Comprehensive Evaluation of GeneLEAD VIII DNA Platform Combined to Deeplex Myc-TB® Assay to Detect in 8 Days Drug Resistance to 13 Antituberculous Drugs and Transmission of Mycobacterium tuberculosis Complex Directly From Clinical Samples, Frontiers in cellular and infection microbiology, 2021 oct., 11:707244.
- Patrick D. Mathews et al, Phylogenetic analysis and characterization of a new parasitic cnidarian (Myxosporea: Myxobolidae) parasitizing skin of the giant mottled eel from the Solomon Islands, Infection, Genetics and Evolution, 2021 oct., 94:104986
- Mathieu Weil et al, Authenticating wild Piper species (peppers) originating from islands in the Indian Ocean on the basis of morphological, genetic and chemical characteristics, Phytochemistry, 2021 oct., 190:112886
- Mona Hoppenrath et al, Molecular phylogeny and morphology of Carinadinium nov. (Dinophyceae, Gonyaulacales), including marine sand-dwelling dinoflagellate species formerly classified within Thecadinium, European Journal of Protistology, 2021 oct., 81:125835
- Simon Begrem et al, Genomic diversity of Serratia proteamaculans and Serratia liquefaciens predominant in seafood products and spoilage potential analyses, International Journal of Food Microbiology, 2021 sept., 354:109326
- Anabel Ordaz-Vázquez et al, Genetic diversity and primary drug resistance transmission in Mycobacterium tuberculosis in southern Mexico, Infection, Genetics and Evolution, 2021 sept., 93:104994
- Maria Prieto-Espinoza et al, Water table fluctuations affect dichloromethane biodegradation in lab-scale aquifers contaminated with organohalides, Water Research, 2021 sept., 203:117530
- François Sipieter et al, Characteristic ERK1/2 signaling dynamics distinguishes necroptosis from apoptosis, iScience, 2021 sept., 24(9):103074
- Nicolas Oury et al, Forensic genetic identification of sharks involved in human attacks, Forensic Science International: Genetics, 2021 sept., 54:101558
- Riccardo Alagna et al, Is the new WHO definition of extensively drug-resistant tuberculosis easy to apply in practice?, The European respiratory journal, 2021 jul., 58(1):2100959
- Amaia Nogales et al, The effects of field inoculation of arbuscular mycorrhizal fungi through rye donor plants on grapevine performance and soil properties, Agriculture, Ecosystems & Environment, 2021 jun; 313:107369
- Rahma Attia El Hili et al, Cytochrome c oxydase I phylogenetic analysis of Haemogregarina parasites (Apicomplexa, Coccidia, Eucoccidiorida, Haemogregarinidae) confirms the presence of three distinct species within the freshwater turtles of Tunisia, Parasitology International, 2021 jun; 82:102306
- Isabelle Simon et al, A Large Insertion Domain in the Rho Factor From a Low G + C, Gram-negative Bacterium is Critical for RNA Binding and Transcription Termination Activity, Journal of Molecular Biology, 2021 may; 433:167060
- Willem Landman et al, First record of Metapolystoma (Monogenea: Polystomatidae) from Boophis tree frogs in Madagascar, with the description of five new species, International Journal for Parasitology: Parasites and Wildlife, 2021 apr; 14:161-178
- Soler et al, Field management of Rotylenchulus reniformis on pineapple combining crop rotation, chemical-mediated induced resistance and endophytic bacterial inoculation, Crop Protection, 2021 march ;141:105446
- Maurine D'Agostino et al, Characterisation of the antifungal effects of a plant-based compound, CIN-102, on the main septal filamentous fungi involved in human pathology, Journal of Global Antimicrobial Resistance, 2021 march, 25:171-180
- Priti Kambli et al, Targeted next generation sequencing directly from sputum for comprehensive genetic information on drug resistant Mycobacterium tuberculosis, Tuberculosis, 2021 march, 127:102051
- Yuka Munakata et al, Composition and functional comparison of vetiver root endophytic microbiota originating from different geographic locations that show antagonistic activity towards Fusarium graminearum, Microbiological Research, 2021 feb; 243:126650
- Oveis Jamialahmadi et al, Exome-Wide Association Study on Alanine Aminotransferase Identifies Sequence Variants in the GPAM and APOE Associated With Fatty Liver Disease, Gastroenterology, 2021 feb
- Stephen Mulero et al, Malacological survey in a bottle of water: A comparative study between manual sampling and environmental DNA metabarcoding approaches, Global Ecology and Conservation, 2021 jan; 25:e01428
- Saliha Ait Yahia et al, NOD1 sensing of house dust mite–derived microbiota promotes allergic experimental asthma, Journal of Allergy and Clinical Immunology, 2021 jan; 148(2):394-406
- Pauline Menelle et al, Biotransformation of guttiferones, Symphonia globulifera metabolites, by Bipolaris cactivora, an endophytic fungus isolated from its leaves, Organic & Biomolecular Chemistry, 2021 jan; 19(6):1378-1385
- Niaz Banaei et al, Novel Assays/Applications for Patients Suspected of Mycobacterial Diseases, Clinics in Laboratory Medicine, 2020 dec; 40(4):535-552,
- Elisabet Alacid et al, Description of two new coexisting parasitoids of blooming dinoflagellates in the Baltic sea: Parvilucifera catillosa sp. nov. and Parvilucifera sp. (Perkinsea, Alveolata), Harmful Algae, 2020 dec; 100:101944
- Priti Kambli & Camilla Rodrigues, Molecular Diagnosis of Drug Resistance in Mycobacterium tuberculosis: Perspectives from a Tuberculosis-Endemic Developing Country, Advances in Molecular Pathology, 2020 nov; 3:87-95
- Thomas Schön et al, What is the role of the EUCAST reference method for MIC testing of the Mycobacterium tuberculosis complex?, Clinical Microbiology and Infection, 2020 nov; 26(11):1453-1455
- Hueso T et al., Impact and consequences of intensive chemotherapy on intestinal barrier and microbiota in acute myeloid leukemia: the role of mucosal strengthening. Gut Microbes. 2020 nov; 12(1):1800897.
- Céline L. Pouille et al. Chicory root flour – A functional food with potential multiple health benefits evaluated in a mice model, Journal of Functional Foods, 2020 Nov; 74:104174
- Luen-Luen Li et al, Impacts of microplastics exposure on mussel (Mytilus edulis) gut microbiota, Science of The Total Environment, 2020 Nov; 745:141018
- Emma Jones et al, Identification of novel risk loci and causal insights for sporadic Creutzfeldt-Jakob disease: a genome-wide association study, The Lancet Neurology, 2020 Oct; 19(10):840-848
- Laura Di Lodovico et al, Anorexia nervosa and gut microbiota: A systematic review and quantitative synthesis of pooled microbiological data, Progress in Neuro-Psychopharmacology and Biological Psychiatry, 2020 Sept; 110114
- Robin Rabier et al, Genetic assessment of a conservation breeding program of the houbara bustard (Chlamydotis undulata undulata) in Morocco, based on pedigree and molecular analyses, Zoo Biology. 2020 Sept; 1– 14
- Marine Le Guyader et al. Successful Leptospira genotyping strategy on DNA extracted from canine biological samples, Journal of Microbiological Methods, 2020 Sept; 176:106007,
- Maxime Branger et al. The complete genome sequence of Mycobacterium bovis Mb3601, a SB0120 spoligotype strain representative of a new clonal group, Infection, Genetics and Evolution, 2020 Aug; 82:104309
- Yaël Escobar et al. Biology, ecology, and impact of Cryptonevra nigritarsis Duda, a potential biological control agent against the giant reed Arundo donax, Biological Control, 2020 Aug; 147:104287
- Julia Mougin et al, Development of a mreB-targeted real-time PCR method for the quantitative detection of Vibrio harveyi in seawater and biofilm from aquaculture systems, Aquaculture, 2020 Aug; 525:735337
- Eleanor S. Click et al, Phylogenetic diversity of Mycobacterium tuberculosis in two geographically distinct locations in Botswana – The Kopanyo Study, Infection, Genetics and Evolution, 2020 Jul; 81:104232
- Matúš Dohál et al, Whole-genome sequencing and Mycobacterium tuberculosis: Challenges in sample preparation and sequencing data analysis, Tuberculosis,2020 Jul; 123:101946
- Cecilia T. Satta et al, Ecological, morphological and molecular characterization of Kryptoperidinium sp. (Dinophyceae) from two Mediterranean coastal shallow lagoons, Harmful Algae,2020 Jul; 97:101855
- Lucas Morinière et al, Clarifying the taxonomy of the causal agent of bacterial leaf spot of lettuce through a polyphasic approach reveals that Xanthomonas cynarae Trébaol et al. 2000 emend. Timilsina et al. 2019 is a later heterotypic synonym of Xanthomonas hortorum Vauterin et al, 1995, Systematic and Applied Microbiology,2020 Jul; 43 (4):126087
- Simon Poirier et al, Large-scale multivariate dataset on the characterization of microbiota diversity, microbial growth dynamics, metabolic spoilage volatilome and sensorial profiles of two industrially produced meat products subjected to changes in lactate concentration and packaging atmosphere, Data in Brief, 2020 jun; 30:105453
- Jean Claude S. Ngabonziza et al. Prevalence and drivers of false-positive rifampicin-resistant Xpert MTB/RIF results: a prospective observational study in Rwanda, The Lancet Microbe, 2020 Jun; 1(2):e74-e83
- Ben Hamou et al, Association of transcription factor 7-like 2 gene (TCF7L2) polymorphisms with stress-related hyperglycaemia (SRH) in intensive care and resulting outcomes: The READIAB study, Diabetes; Metabolism, 2020 Jun; 46(3):243-247
- Imen Hadjadji et al, Unsuspected intraspecific variability in the toxin production, growth and morphology of the dinoflagellate Alexandrium pacificum R.W. Litaker (Group IV) blooming in a South Western Mediterranean marine ecosystem, Annaba Bay (Algeria), Toxicon, 2020 Jun; 180:79-88
- Ehssan Torabi et al, Dissipation of S-metolachlor and butachlor in agricultural soils and responses of bacterial communities: Insights from compound-specific isotope and biomolecular analyses, Journal of Environmental Sciences, 2020 Jun; 92:163-175
- Lamiaa H. Al-Jamea et al, Genetic analysis of TMPRSS6 gene in Saudi female patients with iron deficiency anemia, Hematology/Oncology and Stem Cell Therapy, 2020 May; Available online
- Dieter Weber et al, Demographic history, range size and habitat preferences of the groundwater amphipod Niphargus puteanus (C.L. Koch in Panzer, 1836), Limnologica, 2020 May; 82:125765
- Marion Cordonnier et al, Effects of urbanization–climate interactions on range expansion in the invasive European pavement ant, Basic and Applied Ecology, 2020 May; 44:46-54
- Silva Tafaj et al, Peculiar features of the Mycobacterium tuberculosis population structure in Albania, Infection, Genetics and Evolution, 2020 March; 78:104136
- Sylvine Durand et al, Promiscuity and sex ratio in the terrestrial isopod Armadillidium vulgare and consequences on genetic diversity, Behavioural Processes, 2020 Feb; 171:104030
- Boatema Ofori-Anyinam et al, Comparative genomics shows differences in the electron transport and carbon metabolic pathways of Mycobacterium africanum relative to Mycobacterium tuberculosis and suggests an adaptation to low oxygen tension, Tuberculosis, 2020 Jan; 120: 101899
- Veterinary Parasitology: Regional Studies and Reports, Volume 18, December 2019, Article 100323
- Veterinary Parasitology: Regional Studies and Reports, Volume 18, December 2019, Article 100323
- Ecotoxicology and Environmental Safety, Volume 181, 15 October 2019, Pages 508-517
- Carbohydrate Polymers, Volume 222, 15 October 2019, Article 114999
- Infection, Genetics and Evolution, Volume 72, August 2019, Pages 31-43
- Antiviral Research, Volume 168, August 2019, Pages 114-120
- Infection, Genetics and Evolution, Volume 72, August 2019, Pages 4-9
- Infection, Genetics and Evolution, Volume 72, August 2019, Pages 44-58
- Infection, Genetics and Evolution, Volume 71, July 2019, Pages 159-165
- Biocatalysis and Agricultural Biotechnology, Volume 20, July 2019, Article 101225
- Systematic and Applied Microbiology, Volume 42, Issue 4, July 2019, Pages 448-456
- Biological Control, Volume 134, July 2019, Pages 1-14
- International Journal of Hydrogen Energy, Volume 44, Issue 31, 21 June 2019, Pages 16199-16211
- Acta Tropica, Volume 194, June 2019, Pages 23-29
- Diabetes & Metabolism, In press, corrected proof, Available online 20 May 2019
- Systematic and Applied Microbiology, Volume 42, Issue 3, May 2019, Pages 302-308
- Clinical Microbiology and Infection, Volume 25, Issue 4, April 2019, Pages 403-405
- Heliyon, Volume 5, Issue 4, April 2019, Article e01455
- Veterinary Parasitology: Regional Studies and Reports, Volume 16, April 2019, Article 100280
- Ecological Engineering, Volume 128, March 2019, Pages 66-76
- Systematic and Applied Microbiology, Volume 42, Issue 2, March 2019, Pages 232-239
- Tuberculosis, Volume 115, March 2019, Pages 81-88
- The Lancet Respiratory Medicine, Volume 7, Issue 3, March 2019, Pages 227-238
- Applied Soil Ecology, Volume 135, March 2019, Pages 33-37
- Science of The Total Environment, Volume 651, Part 1, 15 February 2019, Pages 241-249
- Estuarine, Coastal and Shelf Science, Volume 217, 5 February 2019, Pages 218-225
- International Journal of Cardiology, Volume 272, 1 December 2018, Pages 20-25
- Infection, Genetics and Evolution, In press, corrected proof, Available online 26 December 2018
- Systematic and Applied Microbiology, In press, accepted manuscript, Available online 21 December 2018
- Journal of Clinical Tuberculosis and Other Mycobacterial Diseases, Volume 13, December 2018, Pages 17-21
- The Journal of Nutritional Biochemistry, Volume 62, December 2018, Pages 108-122
- Genomics, In press, corrected proof, Available online 15 November 2018
- Cell Metabolism, Volume 28, Issue 5, 6 November 2018, Pages 737-749.e4
- EBioMedicine, Volume 37, November 2018, Pages 410-416
- Systematic and Applied Microbiology, In press, corrected proof, Available online 26 October 2018
- Clinical Nutrition, In press, corrected proof, Available online 9 October 2018
- Cell Metabolism, Volume 28, Issue 4, 2 October 2018, Pages 557-572.e6
- Molecular Phylogenetics and Evolution, Volume 127, October 2018, Pages 758-769
- EBioMedicine, Volume 36, October 2018, Pages 14-15
- Agriculture, Ecosystems & Environment, Volume 265, 1 October 2018, Pages 384-391
- Molecular and Biochemical Parasitology, Volume 225, October 2018, Pages 1-3
- Systematic and Applied Microbiology, Volume 41, Issue 5, September 2018, Pages 452-459
- Science of The Total Environment, Volume 634, 1 September 2018, Pages 1278-1287
- Marine Environmental Research, Volume 140, September 2018, Pages 433-443
- PLoS One. 2018 Aug 3;13(8)
- Journal of Clinical Tuberculosis and Other Mycobacterial Diseases, Volume 12, August 2018, Pages 21-26
- Biological Control, Volume 123, August 2018, Pages 43-52
- Aquatic Botany, Volume 148, August 2018, Pages 25-28
- Systematic and Applied Microbiology, Volume 41, Issue 4, July 2018, Pages 311-323
- Tuberculosis, Volume 111, July 2018, Pages 109-113
- Journal for Nature Conservation, Volume 43, June 2018, Pages 173-178
- Food Research International, Volume 107, May 2018, Pages 451-461
- Tuberculosis, Volume 110, May 2018, Pages 52-55
- Cell Reports, Volume 23, Issue 4, 24 April 2018, Pages 1072-1084
- Journal of Proteomics, Volume 177, 15 April 2018, Pages 148-157
- Chemosphere, Volume 197, April 2018, Pages 123-134
- Agriculture, Ecosystems & Environment, Volume 257, 1 April 2018, Pages 92-102
- EBioMedicine, Volume 30, April 2018, Pages 167-183
- Microorganisms. 2018 Mar 9;6(1)
- Systematic and Applied Microbiology, Volume 41, Issue 2, March 2018, Pages 113-121
- Harmful Algae, Volume 73, March 2018, Pages 58-71
- Chemosphere, Volume 195, March 2018, Pages 190-200
- International Journal of Infectious Diseases, Volume 67, February 2018, Pages 46-51
- European Journal of Protistology, Volume 62, February 2018, Pages 43-55
- Journal of Infection, Volume 76, Issue 1, January 2018, Pages 55-67
- Molecular Phylogenetics and Evolution, Volume 118, January 2018, Pages 122-134
- Soil Biology and Biochemistry, Volume 115, December 2017, Pages 82-91
- Infection, Genetics and Evolution, Volume 55, November 2017, Pages 251-259
- Neurobiology of Aging, Volume 59, November 2017, Pages 220.e1-220.e9
- Molecular Phylogenetics and Evolution, In press, accepted manuscript, Available online 3 October 2017
- Aquaculture, Volume 479, 1 October 2017, Pages 750-758
- Comparative Biochemistry and Physiology Part C: Toxicology & Pharmacology, Vol 199, September 2017, Pages 28-37
- Microbiological Research, Volume 202, September 2017, Pages 11-20
- Soil Biology and Biochemistry, Volume 112, September 2017, Pages 237-247
- Animal Behaviour, Volume 130, August 2017, Pages 251-261
- Molecular Phylogenetics and Evolution, July 2017, Vol 112, Pages 158-173
- Tuberculosis, July 2017, Vol 105, Pages 60-72
- Virus Research, 2017, June
- Environmental Pollution, May 2017, Vol 224, Pages 185-194
- European Journal of Protistology, April 2017, Vol 58, Pages 9-25
- Systematic and Applied Microbiology, April 2017, Vol40, Issue 3, Pages 135-143
- Molecular Phylogenetics and Evolution, April 2017, Vol 109, Pages 430-446
- Estuarine, Coastal and Shelf Science, March 2017, Vol 188, Pages 81-87
- Acta Tropica, March 2017, Vol 167, Pages 196-203
- Comparative Biochemistry and Physiology Part C: Pharmacology, Toxicology and Endocrinology; 2017, June
- Fungal Ecology, Volume 25, February 2017, Pages 14-21
- Journal of Virological Methods, Volume 240, February 2017, Pages 73-77
- International Journal of Food Microbiology, Volume 238, 5 December 2016, Pages 265-273
- Applied Soil Ecology, Volume 108, December 2016, Pages 176-186
- Infection, Genetics and Evolution, Volume 46, December 2016, Pages 94-101
- Molecular Phylogenetics and Evolution, Volume 105, December 2016, Pages 36-49
- Estuarine, Coastal and Shelf Science, Volume 181, 5 November 2016, Pages 256-265
- Infection, Genetics and Evolution, Volume 45, November 2016, Pages 198-204
- Infection, Genetics and Evolution, Volume 45, November 2016, Pages 165-169
- Phytochemistry, Volume 131, November 2016, Pages 92-99
- Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease, Volume 1862, Issue 10, October 2016, Pages 1861
- Biological Control, Volume 101, October 2016, Pages 69-77
- Ticks and Tick-borne Diseases, Volume 7, Issue 6, October 2016, Pages 1256-1264
- Vaccine, Volume 34, Issue 41, 22 September 2016, Pages 5021-5025
- Tuberculosis, Volume 100, September 2016, Pages 72-81
- Biochimie, Volume 127, August 2016, Pages 59-69
- Journal of Virological Methods, Volume 234, August 2016, Pages 101-106
- Journal of Virological Methods, Volume 234, August 2016, Pages 101-106
- The Lancet Infectious Diseases, Volume 16, Issue 8, August 2016, Pages 971-979
- Aquatic Toxicology, Volume 176, July 2016, Pages 64-75
- Brazilian Journal of Microbiology, Volume 47, Issue 3, July–September 2016, Pages 529-530
- Acta Tropica, Volume 158, June 2016, Pages 170-176
- Fungal Genetics and Biology, Volume 91, June 2016, Pages 1-5
- Genomics Data, Volume 8, June 2016, Pages 91-92
- Infection, Genetics and Evolution, Volume 40, June 2016, Pages 8-16
- European Journal of Soil Biology, Volume 74, May–June 2016, Pages 76-80
- Fungal Biology, Volume 120, Issue 5, May 2016, Pages 711-728
- The Lancet Neurology, Volume 15, Issue 5, April 2016, Pages 455-532
- Biological Conservation, Volume 195, March 2016, Pages 279-291
- Physiology & Behavior, Volume 155, 1 March 2016, Pages 17-24
- Systematic and Applied Microbiology, Volume 39, Issue 2, March 2016, Pages 122-131
- Tuberculosis, Volume 97, March 2016, Pages 57-64
- Biochemical Systematics and Ecology, Volume 64, February 2016, Pages 136-141
- Food Microbiology, Volume 53, Part A, February 2016, Pages 41-50
- Veterinary Parasitology, Volume 216, 30 January 2016, Pages 33-37
- International Journal of Biological Macromolecules, Volume 82, January 2016, Pages 653-662
- Forest Ecology and Management, Volume 358, 15 December 2015, Pages 202-211
- Tuberculosis, Volume 95, Issue 6, December 2015, Pages 802-809
- Journal of Comparative Pathology, Volume 153, Issue 4, November 2015, Pages 231-235
- Molecular Phylogenetics and Evolution, Volume 92, November 2015, Pages 1-10
- Virus Research, Volume 208, 2 October 2015, Pages 110-119
- Biochemical Systematics and Ecology, Volume 62, October 2015, Pages 137-141
- Soil Biology and Biochemistry, Volume 89, October 2015, Pages 1-11
- Environmental and Experimental Botany, Volume 116, August 2015, Pages 20-31
- Infection, Genetics and Evolution, Volume 34, August 2015, Pages 298-306
- Acta Tropica, Volume 146, June 2015, Pages 45-52
- Ecological Engineering, Volume 79, June 2015, Pages 113-119
- Genomics Data, Volume 4, June 2015, Pages 22-23
- Protist, Volume 166, Issue 2, May 2015, Pages 234-263
- International Biodeterioration & Biodegradation, Volume 99, April 2015, Pages 55-65
- Systematic and Applied Microbiology, Volume 38, Issue 2, March 2015, Pages 128-134
- Acta Tropica, Volume 142, February 2015, Pages 79-85
- Biochemical Systematics and Ecology, Volume 58, February 2015, Pages 59-63
- Biochemical Systematics and Ecology, Volume 58, February 2015, Pages 242-246
- International Journal of Food Microbiology, Volume 193, 16 January 2015, Pages 82-90
- Biochimie, Volume 107, Part B, December 2014, Pages 367
- Infection, Genetics and Evolution, Volume 28, December 2014, Pages 15-20
- Infection, Genetics and Evolution, Volume 28, December 2014, Pages 715-724
- Infection, Genetics and Evolution, Volume 27, October 2014, Pages 472-480
- Asian Pacific Journal of Tropical Medicine, Volume 7, Supplement 1, September 2014, Pages S212-S216
- Molecular and Biochemical Parasitology, Volume 196, Issue 2, September 2014, Pages 122-125
- Science of the Total Environment, Volume 490, 15 August 2014, Pages 370-378
- Chemosphere, Volume 108, August 2014, Pages 245-250
- Infection, Genetics and Evolution, Volume 26, August 2014, Pages 58-64
- Systematic and Applied Microbiology, Volume 37, Issue 5, July 2014, Pages 368-375
- Virus Research, Volume 186, 24 June 2014, Pages 135-143
- Marine Pollution Bulletin, Volume 83, Issue 1, 15 June 2014, Pages 302-305
- Clinical Microbiology and Infection, Volume 20, Issue 6, June 2014,
- International Journal of Mycobacteriology, Volume 3, Issue 2, June 2014, Pages 108-116
- Molecular and Biochemical Parasitology, Volume 195, Issue 1, June 2014, Pages 30-33
- Veterinary Parasitology, Volume 202, Issues 3–4, 28 May 2014, Pages 171-179
- Infection, Genetics and Evolution, Volume 23, April 2014, Pages 13-19
- Molecular Phylogenetics and Evolution, Volume 73, April 2014, Pages 202-207
- Research in Microbiology, Volume 165, Issue 3, April 2014, Pages 175-189
- Journal of Hospital Infection, Volume 86, Issue 3, March 2014, Pages 188-193
- Soil Biology and Biochemistry, Volume 70, March 2014, Pages 162-165
- Veterinary Parasitology, Volume 199, Issues 3–4, 31 January 2014, Pages 283-288
- European Journal of Soil Biology, Volume 60, January–February 2014, Pages 16-23
- Fungal Biology, Volume 118, Issue 1, January 2014, Pages 12-31
- Protist, Volume 165, Issue 1, January 2014, Pages 81-92
- Fungal Genetics and Biology, Volume 61, December 2013, Pages 80-89
- Water Research, Volume 47, Issue 19, 1 December 2013, Pages 7066-7077
- Journal of Plant Physiology, Volume 170, Issue 17, 15 November 2013, Pages 1536-1540
- Veterinary Parasitology, Volume 197, Issues 1–2, 18 October 2013, Pages 7-12
- Immuno-analyse & Biologie Spécialisée, Volume 28, Issues 5–6, October–December 2013, Pages 297-308
- Molecular Phylogenetics and Evolution, Volume 69, Issue 1, October 2013, Pages 75-82
- Veterinary Microbiology, Volume 166, Issues 1–2, 27 September 2013, Pages 276-280
- Journal de Mycologie Médicale / Journal of Medical Mycology, Volume 23, Issue 3, September 2013, Pages 196-197
- Protist, Volume 164, Issue 5, September 2013, Pages 673-685
- Infection, Genetics and Evolution, Volume 17, July 2013, Pages 16-22
- Infection, Genetics and Evolution, Volume 17, July 2013, Pages 195-201
- Systematic and Applied Microbiology, Volume 36, Issue 5, July 2013, Pages 351-358
- Brain Research, Volume 1517, 23 June 2013, Pages 1-15
- Infection, Genetics and Evolution, Volume 16, June 2013, Pages 362-368
- Systematic and Applied Microbiology, Volume 36, Issue 4, June 2013, Pages 244-251
- Harmful Algae, Volume 25, May 2013, Pages 39-46
- Harmful Algae, Volume 24, April 2013, Pages 65-79
- Molecular Phylogenetics and Evolution, Volume 66, Issue 3, March 2013, Pages 824-832
- Tuberculosis, Volume 93, Issue 2, March 2013, Pages 246-249
- Zoologischer Anzeiger - A Journal of Comparative Zoology, Volume 252, Issue 1, March 2013, Pages 34-40
- Veterinary Parasitology, Volume 192, Issues 1–3, 18 February 2013, Pages 268-272
- Cytokine, Volume 61, Issue 2, February 2013, Pages 602-607
- Journal of Plant Physiology, Volume 170, Issue 2, 15 January 2013, Pages 225-229
- International Dairy Journal, Volume 27, Issues 1–2, December 2012, Pages 13-21
- Journal of Experimental Marine Biology and Ecology, Volumes 432–433, 30 November 2012, Pages 73-82
- Applied Soil Ecology, Volume 62, November 2012, Pages 42-49
- Applied Soil Ecology, Volume 62, November 2012, Pages 142-146
- The Lancet Neurology, Volume 11, Issue 11, November 2012, Pages 931-933
- Journal of Microbiological Methods, Volume 91, Issue 1, October 2012, Pages 65-72
- Marine Pollution Bulletin, Volume 64, Issue 10, October 2012, Pages 2135-2145
- Molecular and Biochemical Parasitology, Volume 185, Issue 2, October 2012, Pages 154-156
- Biochimie, Volume 94, Issue 9, September 2012, Pages 2045-2053
- Deep Sea Research Part I: Oceanographic Research Papers, Volume 67, September 2012, Pages 121-132
- International Journal of Mycobacteriology, Volume 1, Issue 3, September 2012, Pages 161-163
- Journal de Mycologie Médicale / Journal of Medical Mycology, Volume 22, Issue 3, September 2012, Pages 273-274
- Soil Biology and Biochemistry, Volume 51, August 2012, Pages 16-19
- Veterinary Microbiology, Volume 158, Issues 1–2, 6 July 2012, Pages 33-41
- Journal of Cystic Fibrosis, Volume 11, Supplement 1, June 2012, Page S82
- Journal of Cystic Fibrosis, Volume 11, Supplement 1, June 2012, Page S82
- Tuberculosis, Volume 92, Issue 3, May 2012, Pages 264-272
- Journal of Immunological Methods, Volume 378, Issues 1–2, 30 April 2012, Pages 88-94
- International Journal of Food Microbiology, Volume 155, Issue 3, 16 April 2012, Pages 199-210
- Journal of Experimental Marine Biology and Ecology, Volumes 416–417, 15 April 2012, Pages 176-184
- Infection, Genetics and Evolution, Volume 12, Issue 3, April 2012, Pages 549-556
- International Journal of Infectious Diseases, Volume 16, Issue 4, April 2012, Pages e303-e309
- Biological Control, Volume 60, Issue 3, March 2012, Pages 312-320
- Systematic and Applied Microbiology, Volume 35, Issue 2, March 2012, Pages 65-72
- Journal of Proteomics, Volume 75, Issue 2, 21 December 2011, Pages 628-641
- Infection, Genetics and Evolution, Volume 11, Issue 8, December 2011, Pages 1873-1880
- Fungal Biology, Volume 115, Issue 10, October 2011, Pages 1038-1050
- Science of The Total Environment, Volume 409, Issue 20, 15 September 2011, Pages 4489-4495
- Systematic and Applied Microbiology, Volume 34, Issue 5, July 2011, Pages 376-384
- Journal of Virological Methods, Volume 173, Issue 2, May 2011, Pages 320-327
- Experimental Cell Research, Volume 317, Issue 6, 1 April 2011, Pages 886-897
- Pathologie Biologie, Volume 59, Issue 1, February 2011, Pages 19-25
- Journal of Molecular Biology, Volume 405, Issue 2, 14 January 2011, Pages 497-518
- Journal of Plant Physiology, Volume 167, Issue 18, 15 December 2010, Pages 1606-1612
- Harmful Algae, Volume 10, Issue 1, November 2010, Pages 88-97
- Marine Environmental Research, Volume 70, Issue 1, July 2010, Pages 1-12
- Journal of Plant Physiology, Volume 167, Issue 6, 15 April 2010, Pages 480-487
- International Journal of Food Microbiology, Volume 137, Issues 2–3, 28 February 2010, Pages 204-213
- Soil Biology and Biochemistry, Volume 42, Issue 2, February 2010, Pages 244-252
- The International Journal of Biochemistry & Cell Biology, Volume 42, Issue 1, January 2010, Pages 80-89
- Acta Tropica, Volume 112, Issue 3, December 2009, Pages 266-269
- Infection, Genetics and Evolution, Volume 9, Issue 6, December 2009, Pages 1336-1344
- Mycological Research, Volume 113, Issue 12, December 2009, Pages 1351-1364
- Chemosphere, Volume 77, Issue 8, November 2009, Pages 1113-1120
- Plant Physiology and Biochemistry, Volume 47, Issue 8, August 2009, Pages 743-752
- Journal of Plant Physiology, Volume 166, Issue 7, 1 May 2009, Pages 739-749
- Molecular Phylogenetics and Evolution, Volume 50, Issue 3, March 2009, Pages 446-470
- Brain Research, Volume 1249, 16 January 2009, Pages 34-42
- Research in Microbiology, Volume 160, Issue 1, January–February 2009, Pages 63-70
Quality & certifications - High standards and excellence for the benefit of our customers
Very early on, quality has become one of GenoScreen's priorities. This choice ensured that the company’s organizational structure and processes fully meet our customers’ and partners’ requirements. It also means that we comply integrally with all the local and international regulations applicable to GenoScreen’s services.
GenoScreen’s management has committed to guaranteeing:
- The quality and reliability of the services provided,
- The high-performance levels and reliability of the products sold,
- The accuracy of the results provided,
- Long-term customer satisfaction,
- Flexible services that meet the customer’s requirements,
- Continuous improvement of the company’s organizational procedures.
This commitment to quality testifies to the values that structure our activities:
- Customer satisfaction,
- Continuous improvement of the quality of the company’s services and products,
- High performance through innovation.
ISO13485: 2016 certification, Medical devices - Quality management systems - Requirements for regulatory purposes
GenoScreen is ISO13485 certified for the "design, development, manufacture, marketing and distribution of in vitro diagnostic kits. ». This certification completes the ISO9001 certification.
ISO9001 certification : 2015 certification, Quality management system - Requirements
 GenoScreen is ISO9001-certified for the design, development, production and sale of all types of products, services, expert analyses and training courses in genomics and bioinformatics.
GenoScreen is ISO9001-certified for the design, development, production and sale of all types of products, services, expert analyses and training courses in genomics and bioinformatics.
Authorization to handle highly pathogenic microorganisms and toxins (MOT)
 GenoScreen has been authorized by the French drug safety agency to store, process and use highly pathogenic microorganisms and toxins (MOTs). This authorization guarantees that MOTs are handled safely and tracked fully.
GenoScreen has been authorized by the French drug safety agency to store, process and use highly pathogenic microorganisms and toxins (MOTs). This authorization guarantees that MOTs are handled safely and tracked fully.
Accreditation for highly pathogenic microorganisms and toxins (MOTs) by the ANSM (valid until 26/04/2023), in accordance with the French governmental decree #2010-736 dated June 30, 2010.
GCLP: Good Clinical laboratory Practice
 Good Clinical Laboratory Practice (GCLP) is a well-established international quality system for laboratories that analyze clinical trial samples in accordance with Global Good Clinical Practice (GCP) regulations.
Good Clinical Laboratory Practice (GCLP) is a well-established international quality system for laboratories that analyze clinical trial samples in accordance with Global Good Clinical Practice (GCP) regulations.
The requirements of this quality system guarantee patient's rights, preserve their confidentiality and ensure the reliability, quality and integrity of the work and results produced. The evaluation of our Metabiote® protocol: analysis of microbial communities by targeted metagenomics was performed in accordance with the EMA standard (28 February 2012) EMA / INS / GCP / 532137/2010
CIR: 2020 - 2024, Crédit Impôt Recherche (Research Tax Credit)
 GenoScreen benefits from the Research Tax Credit, a measure to support companies' research and development (R&D) activities.
GenoScreen benefits from the Research Tax Credit, a measure to support companies' research and development (R&D) activities.
Oxford Nanopore Technologies, Certified Service Provider

GenoScreen is a certified service provider by Oxford Nanopore Technologies to ensure the highest quality standards for long-read sequencing projects on the GridION platform.
International - Global markets: a leader in genomics
As a market leader for many genomic technologies, GenoScreen’s export activity has developed rapidly. The company now provides its solutions to research groups worldwide.
An international leader
The company sells in over 40 countries.
Turnover breakdown:
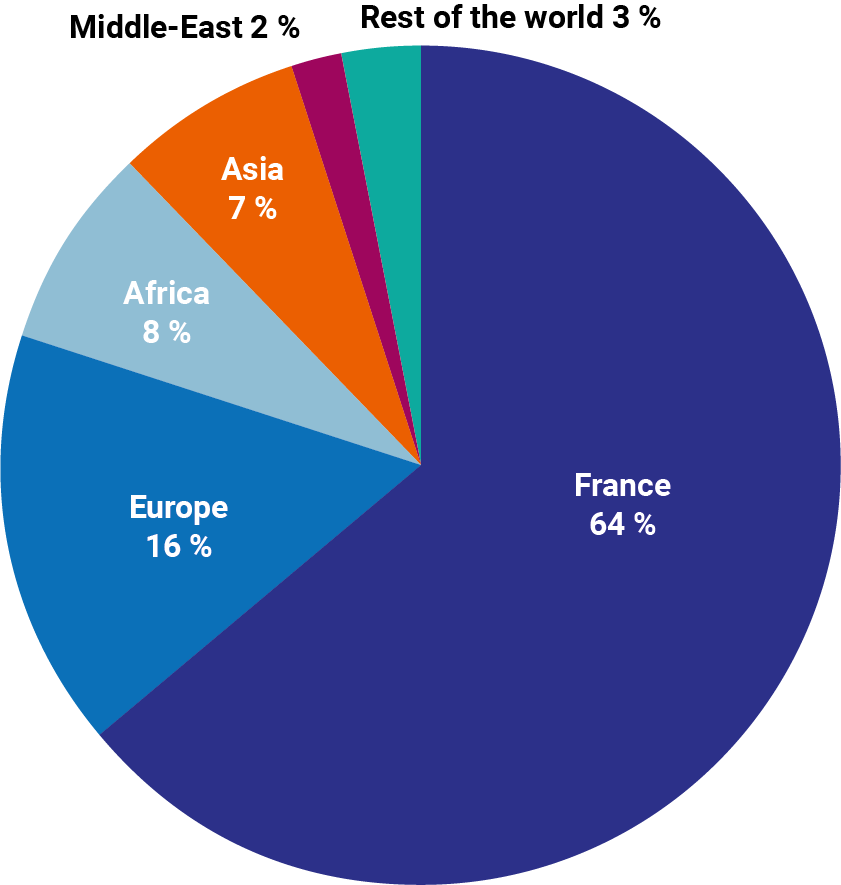
World-class innovations
Some of GenoScreen’s products and solutions have become international benchmarks:
- Deeplex® Myc-TB, all-in-one solution for characterizing Mycobacterium tuberculosis and detecting its antibiotic resistance,
- Deeplex® Myc-Lep, a solution for characterizing Mycobacterium leprae and detecting its antibiotic resistance,
- GenoBiome®, our complete offer for the analysis of microorganism communities (skin with GenoBiome® Skin).
A major player in international research programs
GenoScreen’s R&D teams are frequently involved in world-class research programs. This involvement enables the company to establish and develop close relationships with internationally recognized research groups in the healthcare, environmental and nutrition sectors.
Contact
Address:
GenoScreen,
1 rue du Pr. Calmette,
59000 LILLE, France
Phone numbers:
Reception service: +33(0) 362 263 777
Commercial service : +33(0) 362 263 776
E-mail :
View our Privacy Policy.
Technology platforms - The finest technologies in genomics and bioinformatics
GenoScreen has developed a cutting-edge molecular biology laboratory for the exploitation and the analysis of DNA. Our teams work with a wide range of technologies in order to perform optimized, quality-controlled analyses.
Our laboratory is equipped with cutting-edge technical facilities and the latest generation of automated systems (from Illumina, Applied Biosystems® - Life Technologies, Tecan, Hamilton, Opentrons...)
GenoScreen also has access to a high-security P3 laboratory for handling BSL2 and BSL3 pathogens (Anthrax, BSE, West Nile virus, SARS, tuberculosis, typhus, yellow fever...)
Sanger sequencing
3730XL capillary sequencers (Applied Biosystems®)
High-throughput NGS platforms
Access to all platforms based on Illumina technology (Miseq, Novaseq, etc.)
Third-generation sequencing platform
- GridION (Oxford Nanopore Technologies)
- Sequel IIe (PacBio)
Quantitative PCR and genotyping platform
7900 HT (Applied Biosystem)
Bioinformatics tools
Through its servers, GenoScreen has its own computing power of several hundred logical processors. Through the development of software suites and pipelines, our bioinformaticians have built automated genomic and metagenomic analysis pipelines for robust, rapid and comprehensive analysis, based on our reference software suites:
- Deeplex® software suite for the study of isolated microorganisms,
- Taxonomic classification,
- Functional metagenomics,
- Metatranscriptomics,
- De novo assembly,
- Differential gene expression analysis,
- Variant search (SNPs, Indels, structural variants),
- Phylogeny,
- Search for genes of interest.
Subcategories
GenoScreen
Partners
GenoScreen develops its technologies and know-how in genomics within a dense, well-structured network of partners.
These collaborations co-fund ambitious research programs and jointly develop innovative technologies: an effective and agile strategy that opens up new opportunities for our staff.
Collaborative research - At the heart of international innovation in genomics
GenoScreen is a partner in collaborative research projects that bring together startups, multinationals and public-sector organizations. Our R&D teams provide their knowledge and expertise in the molecular microbiology of isolated agents and complex communities. This allowed our teams to develop our own projects for elaborating innovative products and services.
Our projects are designed to:
- Improve the diagnosis and management of acute/chronic diseases on humans and animals,
- Characterize and monitor microbial biodiversity, with applications in agronomy, agrifood and environment.
The common feature of these projects is the development of molecular tools for the characterization, monitoring and diagnosis of microbial communities. The key objective is to market simple analytical solutions and (ultimately) preventive, corrective or even therapeutic products based on microorganisms.
